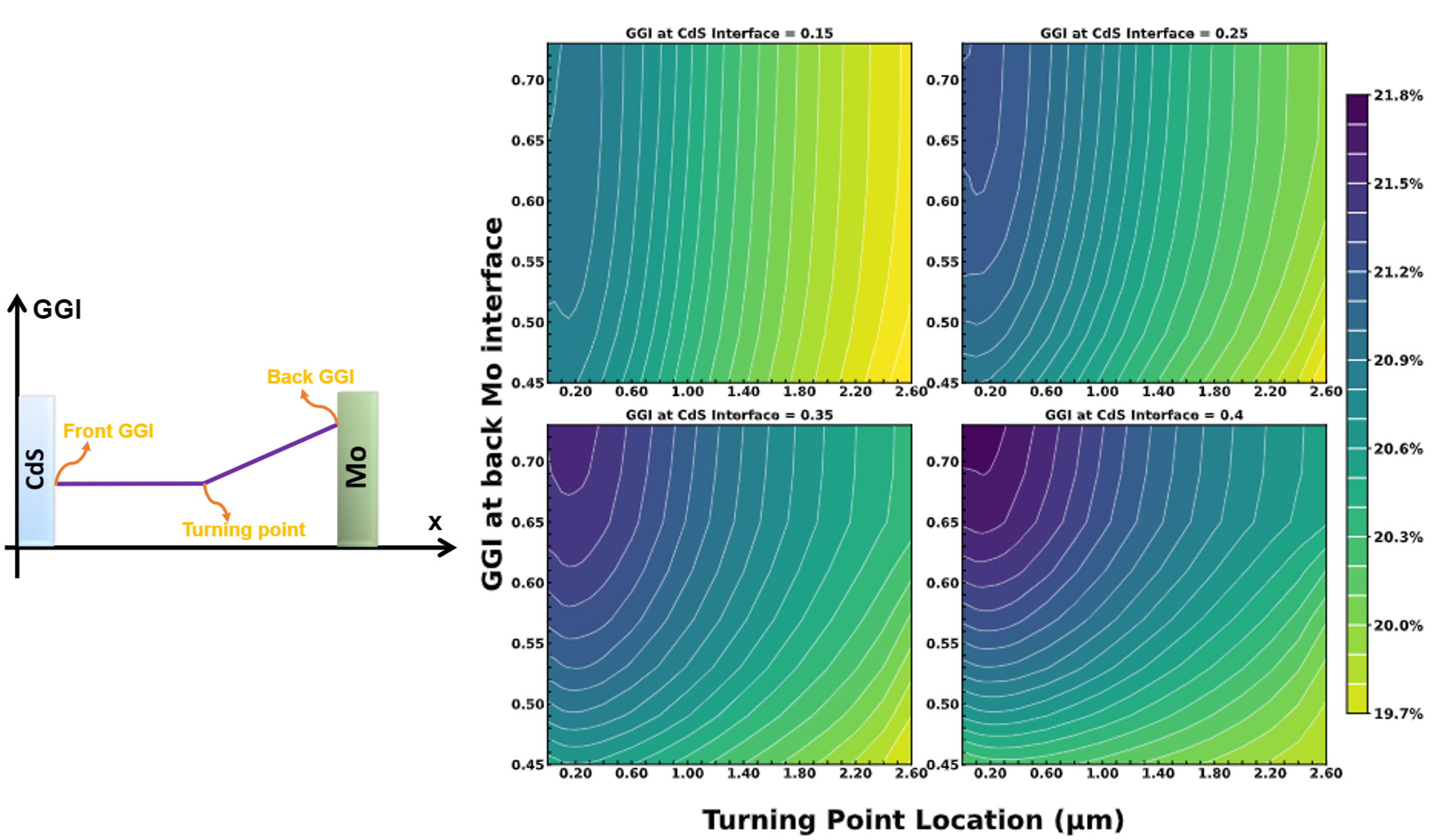
Cu(In,Ga)Se2/CdTeSe Technology Advancement via Fundamental Modeling of Defect/Impurity Interactions [IEEE-PVSC 2021]
Point defects directly impact solar cell device performance by limiting the carrier lifetime. In this work, density functional theory calculations are firstly used to determine the formation energy and diffusion energy barriers of dominant defects in Cu(In,Ga)Se2 and CdSeTe. Next, compositionally constrained therodynamics (CCT) (a C++ package developed in this work) is used to determine equilibrium defect distribution and continuum reaction-diffusion models are developed to analyze the redistribution of defects during manufacturing processes. We estimate defect capture cross sections using a first-principles based approach [CarrierCapture.jl]. These cross sections are combined with our calculated defect profiles and trap energy levels to parameterize a Shockley-Read-Hall (SRH) recombination model, which we implement into a Sentaurus device simulator to predict carrier lifetimes and device performance. In that way, a predictive Technology Computer Aided Design (TCAD) model is built to predict and optimize the performance of Cu(In,Ga)Se2 and CdTeSe solar cells.
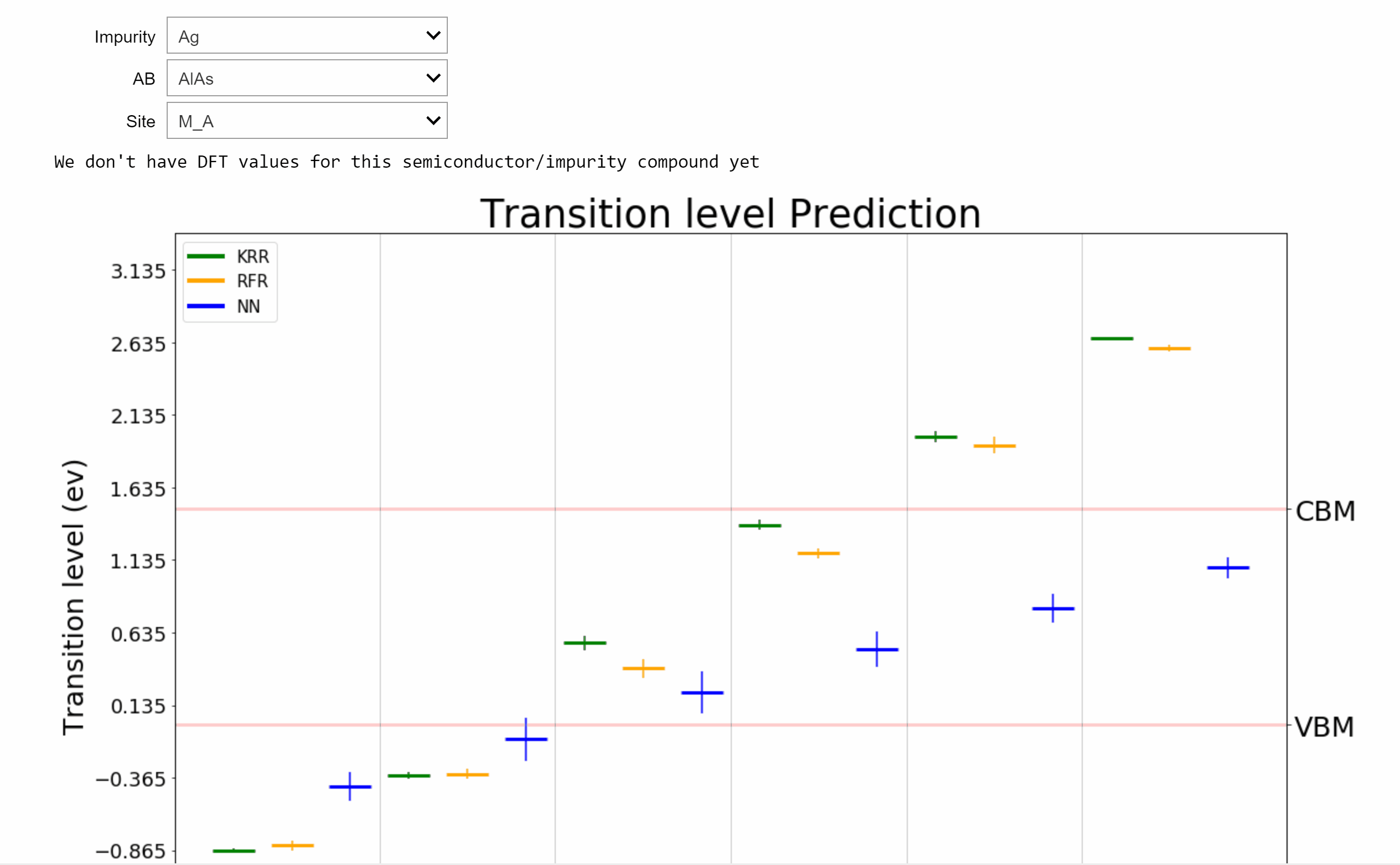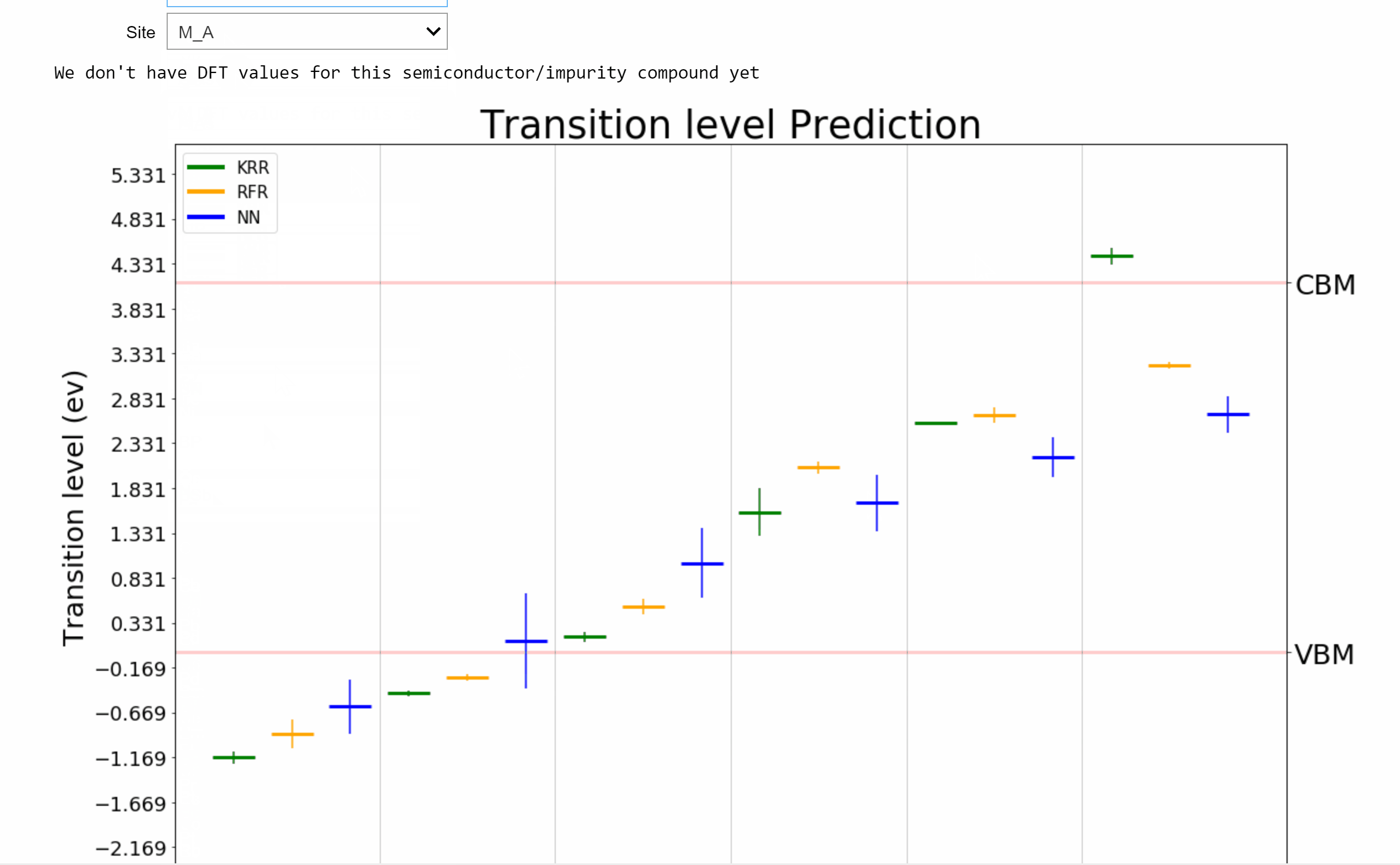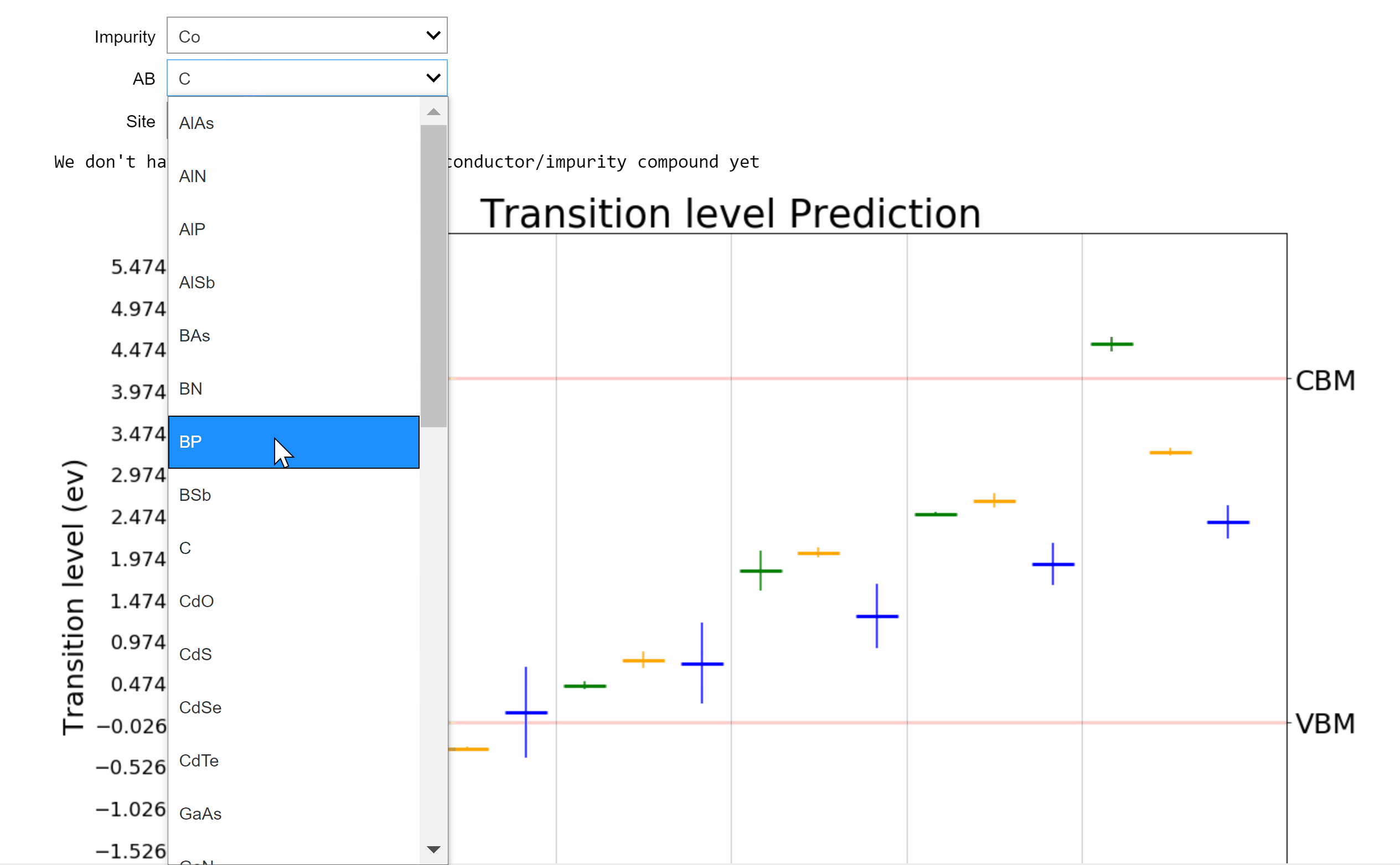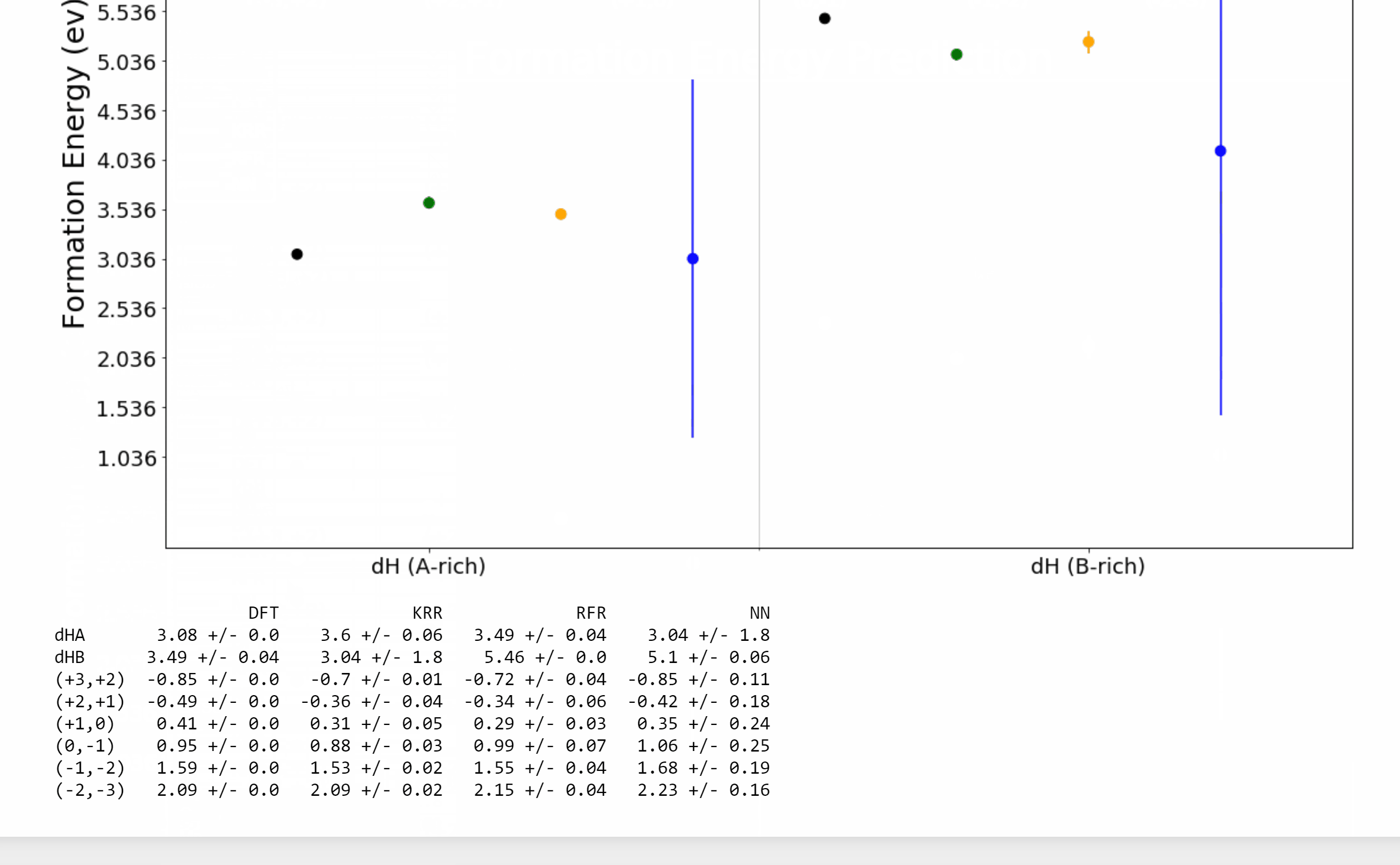
AI-Driven Design of Semiconductors [Patterns 2022]
Point defects or impurities are either naturally present in semiconductors or may be intentionally introduced to tune their electronic and optical properties. The nature of impurity energy levels can strongly influence the performance of a semiconductor in applications ranging from solar cells to photodiodes to infrared sensors to qubits for quantum computing. We develop a framework powered by machine learning (ML) and high-throughput density functional theory (DFT) computations for the prediction and screening of functional impurities in group IV, III-V, and II-VI zinc blende semiconductors. Elements spanning the length and breadth of the periodic table are considered as impurity atoms at the cation, anion, or interstitial sites in supercells of 34 candidate semiconductors, leading to a chemical space of ~12,000 points. Descriptors based on tabulated elemental properties, defect coordination environment, and relevant semiconductor properties are used to train ML regression models for the DFT computed properties, resulting in statistical predictions of the neutral state formation energies and charge transition levels of all possible impurities in the given set of compounds. Kernel ridge regression (KRR), Gaussian process regression (GPR), and neural networks (NN), with appropriate feature selection and hyperparameter optimization, are seen to yield similar predictive performances and meaningful uncertainty estimates. An online tool resulting from this work for predicting and visualizing defect properties in semiconductors is made available on [AI Semiconductor].
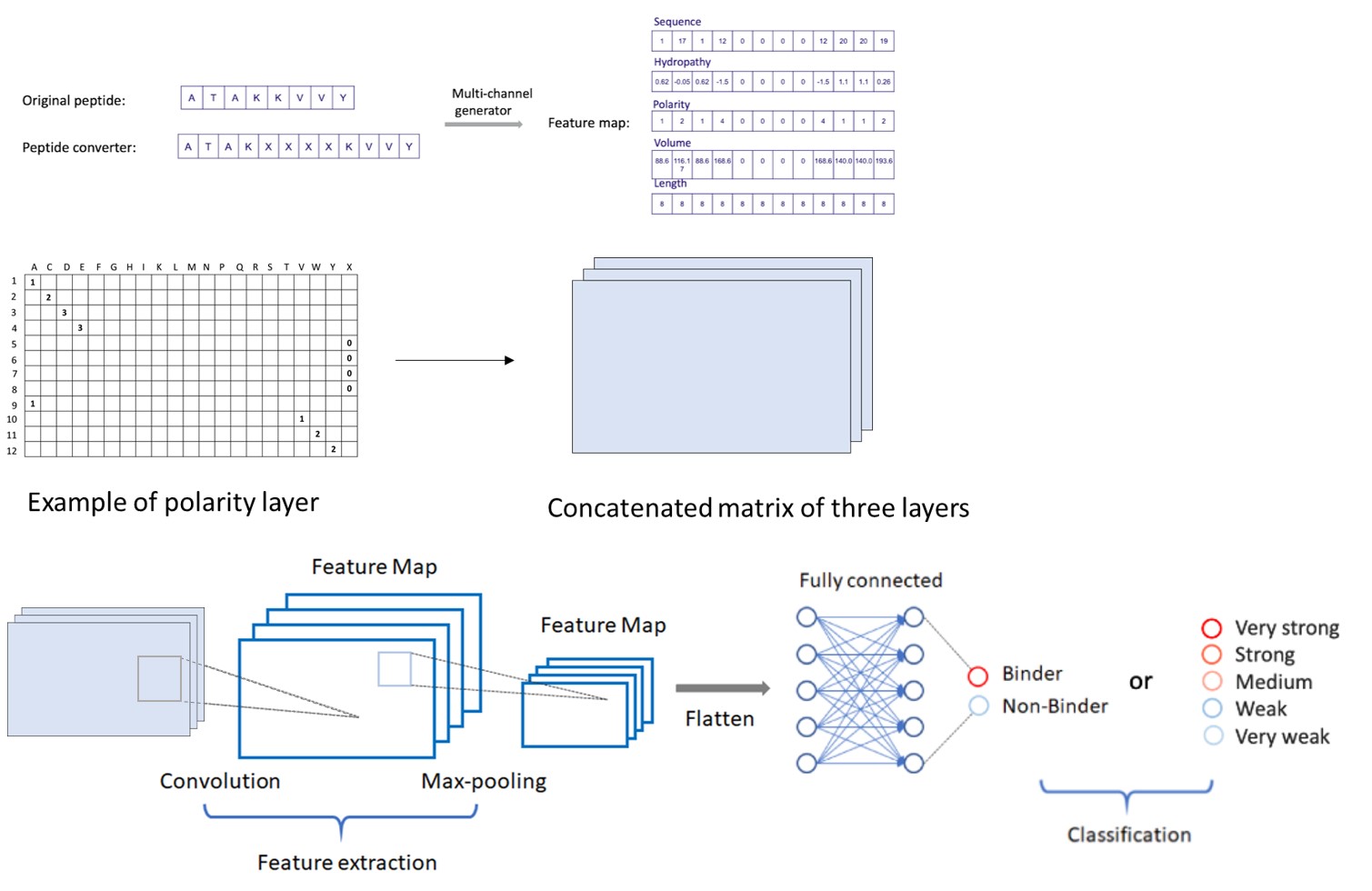
Hyperbinding: a MHC Ligand Prediction Tool Using Convolutional Neutral Network
Hyperbinding is a MHC ligand prediction tool using machine learning approach to generate high-confidence peptides by considering the presentation possibilities of peptides with MHC molecules. MHC-peptide binding prediction plays an important role in cancer immunoengineering, T cell therapy and vaccine design. A simplified immune recognition process involves the binding of a short peptide with antigen presenting cells, where the peptide comes from cancer neoantigen or virus antigen. Higher binding affinity between MHC and the peptide will allow the MHC-peptide complex to be recognized by T cells and thus elicit an immune cascade in eliminating the pathogen. The current release was trained using binding affinity data for HLA-A02:01, which is one of the most abundant alleles in Class I human MHC molecules. Currently, the training and testing support peptides ranging from 8 to 12 amino acids in length, but the model can be re-trained to support more alleles and wider ranges of peptide length. The further application of HyperBinding can be extended into different Class I MHC alleles, and broad categories of disease-related peptides. The prediction tool is available on [Hyperbinding]